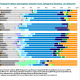
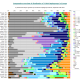
False positives
Bayesian approach which NevGen predictor uses has one bad property, or problem: it assumes that entered haplotype belongs to one of subbranches supported by predictor. So it divides 100% of probability to its supported haplogroups. But this assumption need not be true in every case. No predictor is complete, it does not support every known haplogroup or subbranch. Beside that, user can freely enter any figured-out haplotype, or even random numbers. Even in such cases predictor would give 100% to some of its haplogroups or subbranches, or divide it among several of them. This bad property and biggest NevGen’s weakness till June 11th 2016, is called “False Positive”. Adding new subclades would lower such problems but not solve them completely. For example, R1b level currently supports 168 subclades, but there are some R1b haplotypes that do not belong to any of them. If they were predicted with NevGen prior to June 11th 2016, they would had been FALSE positives, some of predictor’s supported subclades will divide 100% of probability among themselves. The same was with R1a, I1, J1, J2a and all other haplogroups.
Till June 11th 2016, the only way to defend us from such false positives was to look at Fitness score. If it looked too small (for example 12%), it was most probably false positive. In R1b level, most of haplotypes which gave fitness score less than 75 would have been false positives. But, in that time we didn’t had good criteria to tell apart false positives from good predictions.
After first two editions of NevGen, we have had some ideas how to make new and better Fitness scores and by them improve NevGen and ease recognition of false positives. And we implemented them. First, new statistics (calculated from Fitness scores) was added to NevGen Predictor with purpose of giving some estimate that entered haplotype does not belong to any of supported subclades. Now NevGen does not need to distribute 100% of probability to supported subclades, some part of it could be given to “unsupported subclades”, which is big step toward salvation from “False positives” problem. For example, during testing we tested R1b level with several R1a haplotypes with 67 or 111 markers. In all cases, it gave all 100% to “unsupported subclades” and consequently 0% to supported R1b subclades. As was expected. Something must be highlighted here: probability of unsupported subclades is not calculated for haplotypes less than 37 markers long. So, for shorter haplotypes “False positive” problem still exists in its full measure.
Another thing we have done in edition of June 11th 2016 is replacing of older Athey-style Fitness with newer one, based on unbalanced probability score. Since calculation of probability of unsupported subclades is based on Fitness, during mass-testing New Fitness showed much more effective than older one, Athey-style Fitness. So, older one was completely replaced. New Fitness based on probability has considerably lower average value than older one, and it never can be 100%, unlike older one.
Another Fitness added in June 2016 is “Relative Fitness” (Fitness 2), and it is statistics that compares fitness of entered haplotype with average fitness of haplotypes on which was made statistics for given subclade. If haplotype has Relative Fitness greater than 1, it means that it fits better into subclade than average haplotype known to belong to it and used for it’s statistics. It is also calculated only for haplotypes with at least 37 markers.
Because of False positives, but also because of ‘haplotype convergence’ (please see http://isogg.org/wiki/Convergence), our advice to users is to be very cautious and do not blindly trust NevGen Predictor too much, especially in three cases:
1) when number of markers is low (for example 12, or 9).
2) When Fitness score, or Relative Fitness score is low. Low limit for Fitness depends of subclade, for older (like E1a or C2) lower fitness could be good, for example Fitness of 25, but for younger (like R1b DF95) even fitness of 60 could signal false positive. Also, if Relative Fitness (Fitness 2) goes beneath 0.8, it also might signal false positive.
3) when trying to distinguish between very close subclades, like western R1b (under P312).
In no case we guarantee predictions of NevGen nor we take responsibility for any damage from it. NevGen’s predictions are provided “as they are” with no expressed or implied warranty. The authors accept no liability for any damage, in any form, caused by NevGen usage. You use it at your own risk. It has been tested but we are not perfect programmers, nor is data on which NevGen predictor was built.